Identification of clustered microRNAs using an ab initio prediction method.
Posted in Publications, publishedNo Comment
Sewer, A., Paul, N., Landgraf P. et al. Identification of clustered microRNAs using an ab initio prediction method. BMC Bioinformatics, 2005,6, 267.
Abstract
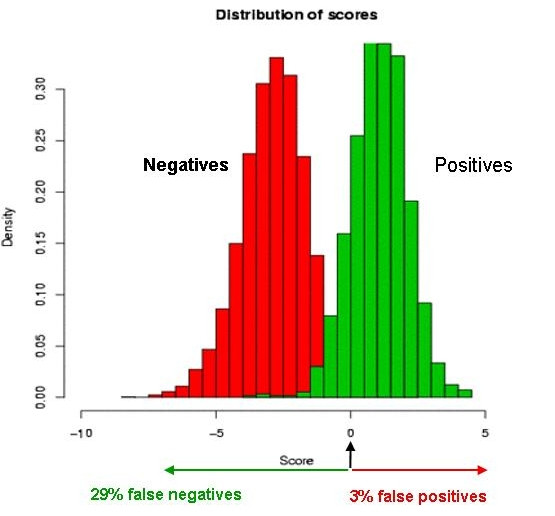
In this work we describe our computational method for miRNA prediction and the results of its application to the discovery of novel mammalian miRNAs. We focus on genomic regions around already known miRNAs, in order to exploit the property that miRNAs are occasionally found in clusters. Starting with the known human, mouse and rat miRNAs we analyze 20 kb of flanking genomic regions for the presence of putative precursor miRNAs (pre-miRNAs). Each genome is analyzed separately, allowing us to study the species-specific identity and genome organization of miRNA loci. We only use cross-species comparisons to make conservative estimates of the number of novel miRNAs. Our ab initio method predicts between fifty and hundred novel pre-miRNAs for each of the considered species. Around 30% of these already have experimental support in a large set of cloned mammalian small RNAs. The validation rate among predicted cases that are conserved in at least one other species is higher, about 60%, and many of them have not been detected by prediction methods that used cross-species comparisons. A large fraction of the experimentally confirmed predictions correspond to an imprinted locus residing on chromosome 14 in human, 12 in mouse and 6 in rat. Our computational tool can be accessed on the world-wide-web.
Download pdf
Supplemental material
Short URL: https://tinyurl.com/y5eah9hy








Comments and Reactions
No comments yet. Why don't you write one?